If you got samflags and want to know their meaning quickly then you can check their meaning interactively following this link. You type in a flag and will get the answer at Piacrd official site.
You can also look up the meaning of samflags in a pdf file sam_output in pdf. Or it looks like this:
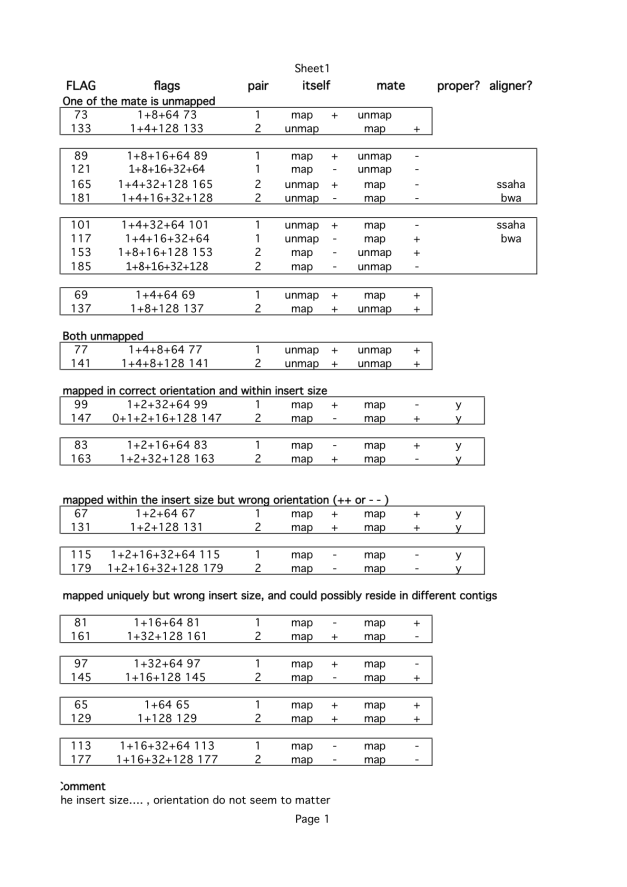 (Provided by WTSI Pathogen Unit). If you do not see the flag you are after, then you can check the picard page or go further below to get actual individual flag components. Or you can just forget about it as an error and check aligners.
(Provided by WTSI Pathogen Unit). If you do not see the flag you are after, then you can check the picard page or go further below to get actual individual flag components. Or you can just forget about it as an error and check aligners.
Please notice positional order of the paired reads “matters” if they are pointing inward or outward.
That is, if you see
A)
1.==> 2.==>
1.<== 2.<==
then the flags are same, since
B)
2.==> 1.==>
2.<== 1.<==
are exactly same to A).
But,
1.==> 2.<==
is not same as
2.<== 1.==>
or vice versa.
This can indicate tandem or episomal duplications or errors. If reads are overlapping, then you see this quite often. (Even experts may not know this. It took me more than 30 minutes convince people.) If you are in doubt, please check all read flags carefully. You may not see them since they can be very rare, which does not mean they do not exist.
It is not easy to identify structural variation (SV) solely based on insert sizes. People unaware of this difficulty think they can just find SVs from their insert sizes: Sorry, but it is not that simple. I was able to find only extremely tandemly repeated gene arrays with this method. Especially it is almost impossible to do this for a draft reference containing many errors.
Here is the flags you should expect after mapping. They are insert size. Note: the orientation of some reads is not defined, and I do not know why. To analyse the flags properly, you may have to order bam files by their name, since you need to know the mapping status of mates. For instance, you need to know both sam flags and positional arrangement of reads. Insert size can help to throw away badly mapped reads but identifying valid SVs is really tough. Or the parasites I am working on may not have very many SVs.
Please check the 1000G paper to find SV detection. The paper A map of human genome variation from population-scale sequencing is available from the Nature web site and is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence to ensure wide distribution.
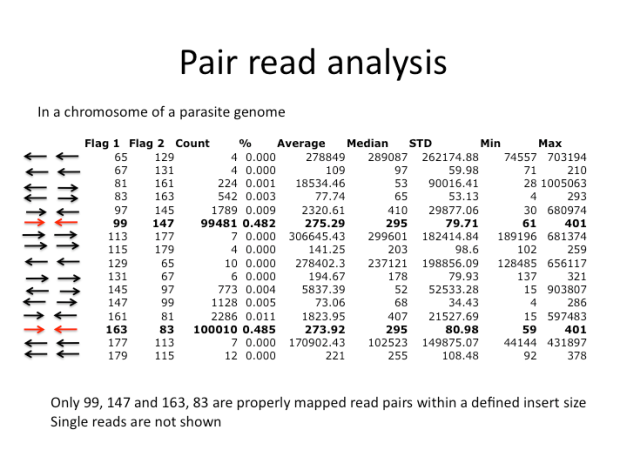 There are many errors in this figure. The direction is not defined in 65 and 129. I need to fix this.
There are many errors in this figure. The direction is not defined in 65 and 129. I need to fix this.
Structural Variant based on Apparent Insert Size:
Small insertion => Decrease
Small deletion => Increase
Large deletion => Increase
Intrachromosomal translocation => Increase
Interchromosomal translocation => Change mate chromosome
Large insertion => Increase or lack of mapped mate
From: Finding Structural Variants in Short Read, Paired-end Sequence Data with R and Bioconductor by Sean Davis
General Information
http://samtools.sourceforge.net/
Other Lang-bindings
Or more generally they are
0x0001 1 the read is paired in sequencing, no matter whether it is mapped in a pair
0x0002 2 the read is mapped in a proper pair (depends on the protocol, normally inferred during alignment)
0x0004 4 the query sequence itself is unmapped
0x0008 8 the mate is unmapped
0x0010 16 strand of the query (0 for forward; 1 for reverse strand)
0x0020 32 strand of the mate
0x0040 64 the read is the first read in a pair
0x0080 128 the read is the second read in a pair
0x0100 256 the alignment is not primary (a read having split hits may have multiple primary alignment records)
========== PrintSamFlag.pl : Usage perl PrintSamFlag.pl input.bam > output
# This program will print out a flag status like this
64706 65 ‘1”64’
61396 69 ‘1”4”64’
66631 73 ‘1”8”64’
130783 77 ‘1”4”8”64’
151001 81 ‘1”16”64’
4225910 83 ‘1”2”16”64’
63678 89 ‘1”8”16”64’
151042 97 ‘1”32”64’
4188825 99 ‘1”2”32”64’
61159 101 ‘1”4”32”64’
63276 113 ‘1”16”32”64’
64707 129 ‘1”128’
66631 133 ‘1”4”128’
61396 137 ‘1”8”128’
130783 141 ‘1”4”8”128’
151042 145 ‘1”16”128’
4200609 147 ‘1”2”16”128’
61159 153 ‘1”8”16”128’
151001 161 ‘1”32”128’
4175327 163 ‘1”2”32”128’
63678 165 ‘1”4”32”128’
63276 177 ‘1”16”32”128’
=======Please copy and paste the following lines and check they are pasted correctly.===========
#!/usr/bin/perl
use strict;
my ($in)=@ARGV;
if ($in=~"\.bam"){
open F1," samtools view $in |" or die;
}
if ($in=~"\.sam"){
open F1,$in or die;
}
my (%flag,%INS,$n1,$n2,@x,$InsS,$P);
while (my $s=<F1>)
{
@x=split '\s+',$s;
++$flag{$x[1]};
++$INS{$x[1]};
#paired
if (SAMflag($x[1])=~"\'2\'"){
#print SAMflag($x[1]) ,"\n";
$InsS+=$x[8];
++$P;
}
}
close F1;
for my $k (sort {$a<=>$b}keys %flag){
print "$flag{$k} $k ",SAMflag($k),"\n";
}
my %TABLE= qw( 0 '0'
1 '1'
2 '2'
3 '1''2'
4 '4'
5 '1''4'
6 '2''4'
7 '1''2''4'
8 '8'
9 '1''8'
10 '2''8'
11 '1''2''8'
12 '4''8'
13 '1''4''8'
14 '2''4''8'
15 '1''2''4''8'
16 '16'
17 '1''16'
18 '2''16'
19 '1''2''16'
20 '4''16'
21 '1''4''16'
22 '2''4''16'
23 '1''2''4''16'
24 '8''16'
25 '1''8''16'
26 '2''8''16'
27 '1''2''8''16'
28 '4''8''16'
29 '1''4''8''16'
30 '2''4''8''16'
31 '1''2''4''8''16'
32 '32'
33 '1''32'
34 '2''32'
35 '1''2''32'
36 '4''32'
37 '1''4''32'
38 '2''4''32'
39 '1''2''4''32'
40 '8''32'
41 '1''8''32'
42 '2''8''32'
43 '1''2''8''32'
44 '4''8''32'
45 '1''4''8''32'
46 '2''4''8''32'
47 '1''2''4''8''32'
48 '16''32'
49 '1''16''32'
50 '2''16''32'
51 '1''2''16''32'
52 '4''16''32'
53 '1''4''16''32'
54 '2''4''16''32'
55 '1''2''4''16''32'
56 '8''16''32'
57 '1''8''16''32'
58 '2''8''16''32'
59 '1''2''8''16''32'
60 '4''8''16''32'
61 '1''4''8''16''32'
62 '2''4''8''16''32'
63 '1''2''4''8''16''32'
64 '64'
65 '1''64'
66 '2''64'
67 '1''2''64'
68 '4''64'
69 '1''4''64'
70 '2''4''64'
71 '1''2''4''64'
72 '8''64'
73 '1''8''64'
74 '2''8''64'
75 '1''2''8''64'
76 '4''8''64'
77 '1''4''8''64'
78 '2''4''8''64'
79 '1''2''4''8''64'
80 '16''64'
81 '1''16''64'
82 '2''16''64'
83 '1''2''16''64'
84 '4''16''64'
85 '1''4''16''64'
86 '2''4''16''64'
87 '1''2''4''16''64'
88 '8''16''64'
89 '1''8''16''64'
90 '2''8''16''64'
91 '1''2''8''16''64'
92 '4''8''16''64'
93 '1''4''8''16''64'
94 '2''4''8''16''64'
95 '1''2''4''8''16''64'
96 '32''64'
97 '1''32''64'
98 '2''32''64'
99 '1''2''32''64'
100 '4''32''64'
101 '1''4''32''64'
102 '2''4''32''64'
103 '1''2''4''32''64'
104 '8''32''64'
105 '1''8''32''64'
106 '2''8''32''64'
107 '1''2''8''32''64'
108 '4''8''32''64'
109 '1''4''8''32''64'
110 '2''4''8''32''64'
111 '1''2''4''8''32''64'
112 '16''32''64'
113 '1''16''32''64'
114 '2''16''32''64'
115 '1''2''16''32''64'
116 '4''16''32''64'
117 '1''4''16''32''64'
118 '2''4''16''32''64'
119 '1''2''4''16''32''64'
120 '8''16''32''64'
121 '1''8''16''32''64'
122 '2''8''16''32''64'
123 '1''2''8''16''32''64'
124 '4''8''16''32''64'
125 '1''4''8''16''32''64'
126 '2''4''8''16''32''64'
127 '1''2''4''8''16''32''64'
128 '128'
129 '1''128'
130 '2''128'
131 '1''2''128'
132 '4''128'
133 '1''4''128'
134 '2''4''128'
135 '1''2''4''128'
136 '8''128'
137 '1''8''128'
138 '2''8''128'
139 '1''2''8''128'
140 '4''8''128'
141 '1''4''8''128'
142 '2''4''8''128'
143 '1''2''4''8''128'
144 '16''128'
145 '1''16''128'
146 '2''16''128'
147 '1''2''16''128'
148 '4''16''128'
149 '1''4''16''128'
150 '2''4''16''128'
151 '1''2''4''16''128'
152 '8''16''128'
153 '1''8''16''128'
154 '2''8''16''128'
155 '1''2''8''16''128'
156 '4''8''16''128'
157 '1''4''8''16''128'
158 '2''4''8''16''128'
159 '1''2''4''8''16''128'
160 '32''128'
161 '1''32''128'
162 '2''32''128'
163 '1''2''32''128'
164 '4''32''128'
165 '1''4''32''128'
166 '2''4''32''128'
167 '1''2''4''32''128'
168 '8''32''128'
169 '1''8''32''128'
170 '2''8''32''128'
171 '1''2''8''32''128'
172 '4''8''32''128'
173 '1''4''8''32''128'
174 '2''4''8''32''128'
175 '1''2''4''8''32''128'
176 '16''32''128'
177 '1''16''32''128'
178 '2''16''32''128'
179 '1''2''16''32''128'
180 '4''16''32''128'
181 '1''4''16''32''128'
182 '2''4''16''32''128'
183 '1''2''4''16''32''128'
184 '8''16''32''128'
185 '1''8''16''32''128'
186 '2''8''16''32''128'
187 '1''2''8''16''32''128'
188 '4''8''16''32''128'
189 '1''4''8''16''32''128'
190 '2''4''8''16''32''128'
191 '1''2''4''8''16''32''128'
192 '64''128'
193 '1''64''128'
194 '2''64''128'
195 '1''2''64''128'
196 '4''64''128'
197 '1''4''64''128'
198 '2''4''64''128'
199 '1''2''4''64''128'
200 '8''64''128'
201 '1''8''64''128'
202 '2''8''64''128'
203 '1''2''8''64''128'
204 '4''8''64''128'
205 '1''4''8''64''128'
206 '2''4''8''64''128'
207 '1''2''4''8''64''128'
208 '16''64''128'
209 '1''16''64''128'
210 '2''16''64''128'
211 '1''2''16''64''128'
212 '4''16''64''128'
213 '1''4''16''64''128'
214 '2''4''16''64''128'
215 '1''2''4''16''64''128'
216 '8''16''64''128'
217 '1''8''16''64''128'
218 '2''8''16''64''128'
219 '1''2''8''16''64''128'
220 '4''8''16''64''128'
221 '1''4''8''16''64''128'
222 '2''4''8''16''64''128'
223 '1''2''4''8''16''64''128'
224 '32''64''128'
225 '1''32''64''128'
226 '2''32''64''128'
227 '1''2''32''64''128'
228 '4''32''64''128'
229 '1''4''32''64''128'
230 '2''4''32''64''128'
231 '1''2''4''32''64''128'
232 '8''32''64''128'
233 '1''8''32''64''128'
234 '2''8''32''64''128'
235 '1''2''8''32''64''128'
236 '4''8''32''64''128'
237 '1''4''8''32''64''128'
238 '2''4''8''32''64''128'
239 '1''2''4''8''32''64''128'
240 '16''32''64''128'
241 '1''16''32''64''128'
242 '2''16''32''64''128'
243 '1''2''16''32''64''128'
244 '4''16''32''64''128'
245 '1''4''16''32''64''128'
246 '2''4''16''32''64''128'
247 '1''2''4''16''32''64''128'
248 '8''16''32''64''128'
249 '1''8''16''32''64''128'
250 '2''8''16''32''64''128'
251 '1''2''8''16''32''64''128'
252 '4''8''16''32''64''128'
253 '1''4''8''16''32''64''128'
254 '2''4''8''16''32''64''128'
255 '1''2''4''8''16''32''64''128');
sub SAMflag{
my $i=shift;
return $TABLE{$i};
}
#======= END ====================
SAMtool bitwise flag: A table of flags with combination of integers in perl format. There is a better way to do this but it is useful, for some.:)
Continue Reading
51.220581
4.399722


